# Open help page for a function
help(mean)
?mean
# Show examples of function usage
example(mean)
# List all built-in datasets
data()Mathew Herrnegger · mathew.herrnegger@boku.ac.at
Institute of Hydrology and Water Management (HyWa) BOKU University Vienna, Austria
LAWI301236 · Distributed Hydrological Modeling with COSERO
Introduction
This document serves as a hands-on introduction to R and RStudio for students working on hydrological modelling and analysis. It covers the R programming foundations required for the Seminar in Surface Hydrology, in particular for working with the CoseRo package in Modules 1–4.
Prerequisites: Basic understanding of hydrology; some programming experience helpful but not required
What This Document Covers
This tutorial is organized into the following main chapters:
- R and RStudio - Overview of the programming language, why it’s useful for hydrology, installation, and the RStudio interface
- Getting Help - How to find answers using built-in help and online resources
- Using LLMs for coding - How to use Claude, ChatGPT etc. in our context
- Basic R Operations - Using R as a calculator and performing hydrological calculations
- Variables and Data Types - Understanding how to store and work with different types of data
- Data Structures - Working with vectors and data frames for organizing hydrological data
- Commenting Code - Best practices for documenting your work
- Working with Data Files - Reading and writing CSV files, Excel files, and using R packages
- Data Manipulation with dplyr - Powerful tools for filtering, transforming, and summarizing data
- Data Visualization - Creating plots to visualize time series, relationships, and distributions
- Practical Example: Enns Catchment - A real-world analysis using Austrian hydrological data
- Programming Essentials - Lists, for-loops, if-else statements, and functions for building efficient workflows
- Summary and Next Steps - Review of key concepts and preparation for the course modules
How to Use This Document
To follow along, copy and paste the code chunks (grey boxes containing R code) directly into RStudio. To copy code, click on the small copy icon in the top-right corner of each code chunk, or manually select the code. You can paste code into a new R script (File > New File > R Script) or directly into the console for immediate execution.
Each section builds on previous concepts, so work through them sequentially. Don’t hesitate to run code multiple times or modify values to better understand how commands work. Making mistakes and debugging are normal parts of learning programming.
R and RStudio
What is R?
R is a programming language and environment for statistical computing and graphics. It was created by Ross Ihaka and Robert Gentleman at the University of Auckland, New Zealand, and first released in 1993 to provide an alternative to the S programming language, which was only available in the commercial S-PLUS package.
Key milestones:
- 1993: R project initiated
- 1995: R released under the GNU General Public License
- 1997: R Core Team formed
- 2000: R version 1.0.0 released
- 2004: CRAN (Comprehensive R Archive Network) established
- 2011: RStudio, Inc. (now Posit) founded, releasing the RStudio IDE
R has become one of the most popular languages for data analysis, statistics, and machine learning.
Why R for Hydrology?
R is useful in hydrology and environmental sciences for:
- Processing and analyzing large datasets
- Analyzing long-term streamflow records
- Handling complex time series data
- Creating visualizations
- Generating hydrographs and other fancy plots
- Creating maps and spatial visualizations
- Implementing statistical models and simulations
- Flood frequency analysis
- Rainfall-runoff modeling
- Reproducible research
- Creating dynamic reports with R Markdown/Quarto
- Sharing code and data for transparency
- Many specialized packages
hydromad,hydroTSM,hydroGOFfor hydrological analysisairGRDelaigue et al. (2023) for hydrological modeling (used in Part 2)- For a somewhat outdated, but comprehensive overview: Slater et al. (2019)
Advantages
- Open-source and free
- Large and active community
- Extensive package ecosystem (~25,000 packages on CRAN)
- Excellent for data visualization
- Supports various data formats (CSV, NetCDF, shapefiles, etc.)
- Strong capabilities in statistical analysis
Considerations
- Learning curve for beginners
- Can be slower than compiled languages for very large datasets
- Some inconsistencies in syntax across packages
What is RStudio?
RStudio is an integrated development environment (IDE) that makes working with R easier and more efficient. For using RStudio, R must also be installed on your computer.
Installation
- Install R: Download from https://cran.r-project.org/
- Install RStudio: Download from https://posit.co/download/rstudio-desktop/
Follow the installation instructions on each website.
RStudio Interface
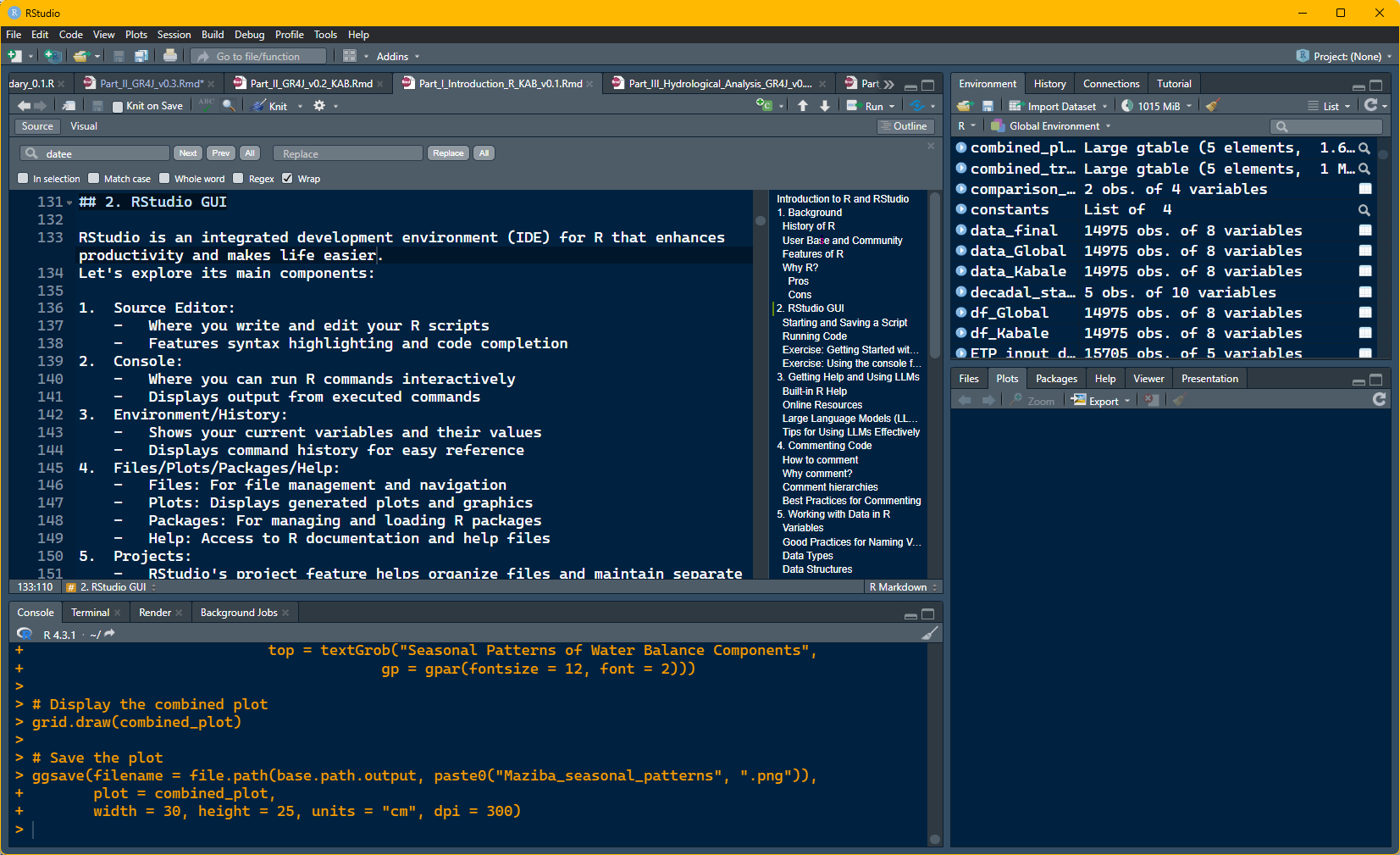
RStudio has four main panels:
- Source Editor (top-left)
- Write and edit R scripts
- Syntax highlighting and code completion
- Console (bottom-left)
- Run R commands interactively
- Displays output from executed commands
- Environment/History (top-right)
- Shows current variables and their values
- Displays command history
- Files/Plots/Packages/Help (bottom-right)
- Files: File management and navigation
- Plots: Displays generated graphics
- Packages: Managing and loading R packages
- Help: Access to R documentation
Starting and Saving a Script
- New script: File > New File > R Script, or press Ctrl+Shift+N (Cmd+Shift+N on Mac)
- Save script: File > Save, or press Ctrl+S (Cmd+S on Mac)
- Choose a meaningful name with a .R extension
Running Code
- Single line: Place cursor on the line and press Ctrl+Enter (Cmd+Enter on Mac)
- Multiple lines: Select the lines and press Ctrl+Enter
- Entire script: Click “Source” button or press Ctrl+Shift+Enter
Exercise: Getting Started
- Open RStudio and create a new R script
- Type:
print('Hello, Hydrology!') - Run the code and observe the output in the console
- Create a variable
flow_rate <- 5.2 - Print the value:
print(flow_rate) - Save the script with a meaningful name
Posit has released Positron, a new free IDE designed for data scientists working with both Python and R, with its second stable desktop release (version 2025.08.0) now available after more than two years of development. Built on the same foundation as Visual Studio Code (Code OSS), Positron provides a cohesive experience for writing code, performing analyses, and exploring data, with native support for plotting and data output across both languages. The IDE includes specialized data science features such as a variable and data frame explorer with interactive filtering and sorting, multi-session consoles for running Python or R code, and integrated notebook support. Posit has emphasized that RStudio is not going away, as it includes over 14 years of R-focused optimizations, and the company remains committed to maintaining and updating RStudio alongside the development of Positron. For those interested in trying Positron, free downloads are available, and migration guides are provided for users transitioning from either VS Code or RStudio.
Getting Help
When learning R or working on projects, you’ll frequently need help. Several resources are available:
Built-in R Help
R provides built-in help functions. These commands open documentation in the Help panel of RStudio, showing function descriptions, parameters, and usage examples.
In RStudio, press F1 while the cursor is on a function name to open its help page.
Online Resources
- Google: Often the best starting point. Include “R” in your search query
- Example: “how to read csv file in R”
- Search for exact error messages in quotes
- Stack Overflow: https://stackoverflow.com/questions/tagged/r
- Community Q&A platform
- R-bloggers: https://www.r-bloggers.com/
- R tutorials and news
- RStudio Community: https://community.rstudio.com/
Using Large Language Models (LLMs)
LLMs like ChatGPT, Claude, or GitHub Copilot are game changers and can be powerful tools for coding assistance. However, they should complement, not replace, your own learning.
Effective Use of LLMs
- Be specific: Clearly state your problem with relevant details
- Provide context: Briefly explain your task or project
- Share code: Include relevant code snippets
- Verify solutions: Always test and understand the code provided
- Ask for explanations: Request comments or reasoning if unclear
- Iterate: Ask follow-up questions if needed
- Learn actively: Use LLMs as a learning tool, not just for solutions
Example 1: Code Generation for Hydrological Analysis
Task: You need to calculate the runoff coefficient (ratio of runoff to precipitation) for multiple years. The runoff coefficient is a dimensionless value indicating the proportion of precipitation that becomes runoff.
Prompt to LLM:
I'm working on a hydrological analysis in R. I have two vectors:
- annual_precip: annual precipitation values in mm (e.g., c(1200, 1100, 1300))
- annual_runoff: annual runoff values in mm (e.g., c(400, 350, 450))
Can you write an R function that calculates the runoff coefficient for each year
and returns a data frame with years, precipitation, runoff, and runoff coefficient?
The function should handle cases where vectors have different lengths.LLM Response (example):
calculate_runoff_coefficient <- function(precip, runoff, years = NULL) {
# Check if vectors have same length
if (length(precip) != length(runoff)) {
stop("Precipitation and runoff vectors must have the same length")
}
# Calculate runoff coefficient
runoff_coeff <- runoff / precip
# Create result data frame
result <- data.frame(
Year = years,
Precipitation_mm = precip,
Runoff_mm = runoff,
Runoff_Coefficient = runoff_coeff
)
return(result)
}
# Example usage
annual_precip <- c(1200, 1100, 1300, 980, 1250)
annual_runoff <- c(400, 350, 450, 300, 420)
years <- 2018:2022
result <- calculate_runoff_coefficient(annual_precip, annual_runoff, years)
print(result) Year Precipitation_mm Runoff_mm Runoff_Coefficient
1 2018 1200 400 0.3333333
2 2019 1100 350 0.3181818
3 2020 1300 450 0.3461538
4 2021 980 300 0.3061224
5 2022 1250 420 0.3360000Key learning points:
- Error checking with
ifstatements andstop() - Creating data frames with multiple columns
- Function documentation through comments
Example 2: Debugging Code with LLM
Scenario: You wrote code to calculate monthly mean discharge from your data, but it’s producing an error.
Your buggy code:
# This code has bugs!
library(lubridate)
discharge_data <- data.frame(
Date = as.Date(c("2020-01-15", "2020-01-20", "2020-02-10",
"2020-02-25", "2020-03-05")),
Discharge = c(15.2, 18.3, 22.1, 19.5, 16.8)
)
# Extract month and calculate monthly means
discharge_data$Month <- month(Date) # BUG HERE!
monthly_mean <- aggregate(Discharge ~ Month,
data = discharge_data,
FUN = mean)
print(monthly_mean)Error message may be something like this:
Error in `as.POSIXlt.default()`:
! do not know how to convert 'x' to class “POSIXlt”
Prompt to LLM:
I'm trying to calculate monthly mean discharge from a data frame in R,
but I'm getting an error. Here's my code and the error message:
[paste your code and error here]
What's wrong and how can I fix it?LLM Response (example):
# Fixed code with explanation
library(lubridate)
discharge_data <- data.frame(
Date = as.Date(c("2020-01-15", "2020-01-20", "2020-02-10",
"2020-02-25", "2020-03-05")),
Discharge = c(15.2, 18.3, 22.1, 19.5, 16.8)
)
# FIX: Reference the data frame column, not just the column name
discharge_data$Month <- month(discharge_data$Date) # CORRECTED!
# Now calculate monthly means
monthly_mean <- aggregate(Discharge ~ Month,
data = discharge_data,
FUN = mean)
print(monthly_mean) Month Discharge
1 1 16.75
2 2 20.80
3 3 16.80Explanation from LLM may be:
“The error occurs on this line: discharge_data$Month <- month(Date)
The problem is that Date is referenced without specifying which data frame it belongs to. R is looking for a variable called Date in the global environment, but it only exists as a column inside discharge_data.
Solution:
Reference the column using discharge_data$Date instead of just Date:
discharge_data$Month <- month(discharge_data$Date) # CorrectThis is a very common mistake! When working with data frame columns, you need to tell R which data frame they come from using the $ operator or by working within specific contexts (like inside dplyr pipes or with() statements).”
Key learning points:
- Understanding R’s scoping: Variables vs. data frame columns
- The importance of referencing data frame columns correctly with
$ - Reading error messages: “object ‘X’ not found” usually means a typo or scoping issue
- Common beginner mistake that even experienced programmers make sometimes!
DO:
- Use LLMs to understand concepts and get starting code
- Ask for explanations of generated code
- Verify all code by running it yourself
- Learn from the patterns and techniques shown
DON’T:
- Blindly copy-paste without understanding
- Skip error checking and validation
- Assume the first solution is optimal
- Use LLM code in production without testing
Remember: LLMs can make mistakes. Always test code and understand what it does before using it in your analysis.
Basic R Operations
R as a Calculator
R can perform basic mathematical operations. Simply type expressions into the console or script, and R evaluates them following standard mathematical order of operations.
# Addition
5 + 3
# Subtraction
10 - 4
# Multiplication
6 * 7
# Division
20 / 4
# Exponentiation
2^3
# Square root
sqrt(16)
# Order of operations
(5 + 3) * 2Hydrological Calculations
These examples demonstrate “typical” unit conversions and hydraulic calculations used in hydrology. Manning’s equation calculates flow velocity in open channels based on roughness, hydraulic radius, and slope.
# Convert flow from m³/s to L/s
flow_m3s <- 5.3
flow_Ls <- flow_m3s * 1000
print(flow_Ls)
# Calculate the area of a circular pipe (in m²) with diameter 0.4 m
diameter <- 0.4
area <- pi * (diameter/2)^2
print(area)
# Calculate Manning's equation for flow velocity
# V = (1/n) * R^(2/3) * S^(1/2)
# Where:
# V = flow velocity (m/s)
# n = Manning's roughness coefficient
# R = hydraulic radius (m)
# S = channel slope (m/m)
n <- 0.03 # Manning's coefficient
R <- 0.6 # Hydraulic radius (m)
S <- 0.002 # Slope (m/m)
velocity <- (1/n) * (R)^(2/3) * sqrt(S)
print(paste("Flow velocity:", round(velocity, 2), "m/s"))Variables and Data Types
Variables
Variables are containers that store data values in computer memory. They are fundamental to programming because they allow you to:
- Store calculation results for later use
- Make code more readable by using descriptive names
- Perform operations on stored values
- Update values as your analysis progresses
Think of variables like labeled boxes: you put data in a box and give it a meaningful name so you can find and use it later.
Variable Assignment
In R, use the <- operator to assign values to variables (though = also works, <- is more common in R programming). The following example calculates annual runoff depth from monthly discharge data.
# Simple variable assignment
x <- 10
y <- 5
z <- x + y
print(z)
# Variables can be reassigned (updated)
x <- 20
z <- x + y # z is now 25
print(z)
# Hydrological example: Calculate annual runoff depth
# Given monthly flows in m³/s for the Schladming catchment
# Define monthly flows (m³/s)
monthly_flows <- c(5, 8, 15, 25, 40, 30, 20, 12, 8, 6, 4, 3)
# Calculate annual runoff volume (m³)
days_per_month <- c(31, 28, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31)
annual_volume <- sum(monthly_flows * 3600 * 24 * days_per_month)
# Catchment area (km²)
catchment_area <- 648.8 # Enns catchment above Schladming
# Calculate annual runoff depth (mm)
# 1 m³ = 1000 L = 1000 dm³
# 1 km² = 1,000,000 m²
# 1 mm = 1 L/m²
annual_runoff_depth <- (annual_volume * 1000) / (catchment_area * 1e6)
print(paste("Annual runoff depth:", round(annual_runoff_depth, 1), "mm"))Why Use <- Instead of =?
Both work for assignment, but <- is preferred in R for several reasons:
- Clarity: Makes it clear you’re assigning a value (not testing equality)
- Directionality: Shows data flows from right to left
- Convention: Standard in R community and style guides
- Avoiding confusion:
=is also used for function arguments
# Both work for assignment
discharge1 <- 15.3
discharge2 = 15.3
# But in function calls, use = for arguments
mean_discharge <- mean(c(10, 15, 20), na.rm = TRUE)- Use descriptive names:
annual_rainfallinstead ofar - Use lowercase with underscores (snake_case):
river_discharge - Avoid function names: Don’t use
c,mean,sum, etc. - Start with a letter:
station_1not1station - Be consistent: Use the same style throughout your code
- Use abbreviations sparingly: Ensure they’re still clear
Data Types
Data types define what kind of information a variable holds and what operations can be performed on it. R has several basic data types, each suited for different purposes. Understanding data types is crucial because:
- Different types support different operations (you can’t add text!)
- Functions expect specific data types as inputs
- Incorrect types are a common source of errors
- Type conversions may be needed when reading data
Numeric
Decimal values (floating-point numbers) for continuous measurements. This is the default type for numbers in R.
water_depth <- 3.7 # meters
temperature <- 15.3 # degrees Celsius
discharge <- 23.456 # m³/s
# Mathematical operations work on numeric data
total_depth <- water_depth + 1.3
# check class
print(class(water_depth))[1] "numeric"# Numeric values can have decimals
print(water_depth)[1] 3.7print(temperature)[1] 15.3print(total_depth)[1] 5When to use: Measurements like discharge, precipitation, temperature, concentrations, water levels, etc.
Integer
Whole numbers for count data. In R, you need to explicitly specify integers with the L suffix.
num_sampling_sites <- 12L # The 'L' tells R it's an integer
year <- 2023L
num_days <- 365L
# Without 'L', R treats it as numeric
not_an_integer <- 12
print(class(num_sampling_sites))[1] "integer"print(class(not_an_integer)) # "numeric", not "integer"[1] "numeric"# Integers use less memory (rarely matters in practice)
print(object.size(12))56 bytesprint(object.size(12L))56 bytesWhen to use: Counts (number of stations, years, samples), indices, or when you need to ensure whole numbers.
Character
Text data (strings) for labels, names, categories, and text information.
river_name <- "Enns"
station_name <- "Schladming"
unit <- "m³/s"
# print class
print(class(river_name))[1] "character"# Use quotes for character data
print(river_name)[1] "Enns"# Combine text with paste() or paste0()
full_name <- paste(river_name, "-", station_name)
print(full_name)[1] "Enns - Schladming"# !! You can't do math with characters !!
# This will give an error: river_name + 5When to use: Station names, river names, dates as text, categories, file paths, labels in plots.
Logical
TRUE or FALSE values (also called Boolean) for conditional operations and filtering.
is_flooding <- FALSE
high_flow <- TRUE
data_quality_ok <- TRUE
print(class(is_flooding))
# Logical values result from comparisons
discharge <- 25.3
is_high_discharge <- discharge > 20
print(is_high_discharge) # TRUE
# Useful for filtering
temperatures <- c(5, 15, 25, 10, 20)
above_threshold <- temperatures > 15
print(above_threshold) # FALSE FALSE TRUE FALSE TRUE
# Select only values above threshold
hot_days <- temperatures[above_threshold]
print(hot_days) # 25 20When to use: Conditional logic, filtering data, flags for data quality, controlling if-else statements.
Type Conversion
Sometimes you need to convert between data types:
# Convert character to numeric
year_text <- "2023"
year_numeric <- as.numeric(year_text)
print(class(year_numeric))
# Convert numeric to integer
discharge_numeric <- 15.7
discharge_integer <- as.integer(discharge_numeric) # Becomes 15
print(discharge_integer)
# Convert to character
flow <- 23.5
flow_text <- as.character(flow)
print(flow_text) # "23.5"
# Convert to logical
values <- c(0, 1, 2, 0)
logical_values <- as.logical(values) # 0 becomes FALSE, others TRUE
print(logical_values)Checking Data Types
Use the class() function to verify what type of data a variable contains. This is helpful for debugging and ensuring operations are appropriate for the data type.
# Check the type of a variable
class(water_depth) # "numeric"
class(river_name) # "character"
class(is_flooding) # "logical"
# Alternative functions for type checking
is.numeric(water_depth) # TRUE
is.character(river_name) # TRUE
is.logical(is_flooding) # TRUE
# Check structure of complex variables
str(water_depth) # num 3.7
str(river_name) # chr "Enns"Common Data Type Issues in Hydrology
# Issue 1: Dates imported as character
date_text <- "2023-05-15"
class(date_text) # "character"
# Solution: Convert to Date
date_proper <- as.Date(date_text)
class(date_proper) # "Date"
# Issue 2: Numbers imported as character (e.g., with decimal comma)
discharge_text <- "15,3" # European decimal format
# This won't work: as.numeric(discharge_text) gives NA
# Solution: Replace comma with period first
discharge_fixed <- gsub(",", ".", discharge_text)
discharge_num <- as.numeric(discharge_fixed)
# Issue 3: Missing values coded as text
flow_data <- c("5.2", "6.8", "-999", "7.1") # -999 = missing
flow_numeric <- as.numeric(flow_data) # Works but -999 is a number
flow_numeric[flow_numeric == -999] <- NA # Replace with NAData Structures
Data structures are ways of organizing and storing data in R. Choosing the right data structure is important for:
- Efficiency: Some operations are faster with certain structures
- Clarity: The right structure makes your code easier to understand
- Functionality: Different structures support different operations
R has several data structures, but for hydrology the most important are vectors (for single variables) and data frames (for datasets with multiple variables). Other structures include lists, matrices, and arrays, which we’ll mention briefly but won’t cover in depth.
Vectors
Vectors are one-dimensional arrays that hold data of the same type (all numeric, all character, etc.). They are created using the c() function (short for “combine” or “concatenate”) and are fundamental for storing sequences of measurements like time series data.
Why vectors matter in hydrology:
- Store time series (daily discharge, precipitation, temperature)
- Perform element-wise calculations (e.g., unit conversions on all values at once)
- Use vectorized operations for efficiency (R is optimized for vector operations)
- Essential building blocks for data frames
# Create a numeric vector of daily rainfall
daily_rainfall <- c(5.2, 0, 12.5, 8.7, 6.3)
print(daily_rainfall)
# Length of vector
print(paste("Number of days:", length(daily_rainfall)))
# Create a character vector of river names
river_names <- c("Enns", "Mur", "Drau", "Inn", "Salzach")
print(river_names)
# Create a sequence of years
years <- seq(2018, 2024, by = 1)
print(years)
# Alternative way to create sequences (shortcut)
years_alt <- 2018:2024
print(years_alt)
# Create sequences with specific length
months <- seq(1, 12, length.out = 12)
print(months)
# Perform operations on vectors (vectorized operations!)
mean_rainfall <- mean(daily_rainfall)
print(paste("Mean daily rainfall:", mean_rainfall, "mm"))
# All values are processed at once (vectorization)
rainfall_mm_to_cm <- daily_rainfall / 10
print(rainfall_mm_to_cm)
# Element-wise operations between vectors
rainfall_week1 <- c(5.2, 0, 12.5, 8.7, 6.3, 4.1, 9.8)
rainfall_week2 <- c(3.1, 7.2, 0, 5.5, 11.2, 8.8, 2.9)
total_rainfall <- rainfall_week1 + rainfall_week2
print(total_rainfall)Accessing Vector Elements
Use square brackets [] with an index to extract specific elements. R uses 1-based indexing (the first element is at position 1, unlike Python which starts at 0).
# Access single element (R uses 1-based indexing)
print(daily_rainfall[3]) # Third element
# Access multiple elements
print(daily_rainfall[1:3]) # First three elements
# Access specific positions
print(daily_rainfall[c(1, 3, 5)]) # 1st, 3rd, and 5th elements
# Negative indexing (exclude elements)
print(daily_rainfall[-1]) # All except first element
print(daily_rainfall[-c(1, 2)]) # All except first two
# Access elements by condition (logical indexing)
high_rainfall <- daily_rainfall[daily_rainfall > 8]
print(high_rainfall)
# Logical indexing is very powerful for filtering
discharge <- c(5, 15, 25, 10, 30, 8)
high_flow <- discharge > 20
print(high_flow) # TRUE/FALSE vector
print(discharge[high_flow]) # Only values > 20
# Replace specific values
discharge_modified <- discharge
discharge_modified[discharge_modified > 20] <- 20 # Cap at 20
print(discharge_modified)Vector Operations and Functions
R has many built-in functions for working with vectors:
flow_data <- c(5.2, 8.7, 15.3, 12.1, 9.8, 6.5, 4.2)
# Summary statistics
mean(flow_data) # Average
median(flow_data) # Median
sd(flow_data) # Standard deviation
min(flow_data) # Minimum
max(flow_data) # Maximum
range(flow_data) # Min and max
sum(flow_data) # Sum of all values
length(flow_data) # Number of elements
# Quantiles
quantile(flow_data, probs = c(0.25, 0.50, 0.75))
# Sorting
sort(flow_data) # Ascending order
sort(flow_data, decreasing = TRUE) # Descending order
# Finding positions
which.max(flow_data) # Position of maximum value
which.min(flow_data) # Position of minimum value
which(flow_data > 10) # Positions where condition is TRUE
# Rounding
round(flow_data, 1) # Round to 1 decimal place
floor(flow_data) # Round down
ceiling(flow_data) # Round up
# Cumulative operations
cumsum(flow_data) # Cumulative sum
cumprod(flow_data) # Cumulative productData Frames
Data frames are the most commonly used data structure in R for storing tabular data (like spreadsheets or database tables). They are collections of vectors of equal length, where each vector becomes a column.
Key characteristics:
- Rows represent observations (e.g., daily measurements, gauging stations)
- Columns represent variables (e.g., discharge, temperature, precipitation)
- Columns can have different types (numeric, character, logical, etc.)
- Similar to Excel/CSV files but more powerful and flexible
Why data frames matter in hydrology:
- Store complete datasets with multiple variables (discharge, precip, temp, etc.)
- Each row is typically a time point or station
- Easy to filter, subset, and analyze
- Most R functions expect data frames as input
- Natural way to work with time series and spatial data
Creating and Accessing Data Frames
# Create a data frame with river data
# Source: https://wasser.umweltbundesamt.at/hydjb/search/search.xhtml
# Using most downstream discharge station
river_data <- data.frame(
Name = c("Enns", "Mur", "Drau", "Inn", "Salzach"),
Area_km2 = c(5915.4, 9769.9, 10968.0, 25520.0, 6165.4),
Discharge_m3s = c(191, 138, 271, 724, 237)
)
print(river_data)
# Access a specific column using $
print(river_data$Area_km2)
# Access by column name with brackets
print(river_data[, "Name"])
# Access specific rows
print(river_data[1:3, ]) # First three rows
# Access specific cells [row, column]
print(river_data[2, 3]) # Row 2, Column 3
# Calculate mean discharge
mean_discharge <- mean(river_data$Discharge_m3s)
print(paste("Mean discharge:", round(mean_discharge, 1), "m³/s"))
# View structure of data frame
str(river_data)
# Get dimensions
nrow(river_data) # Number of rows
ncol(river_data) # Number of columns
dim(river_data) # Both dimensions
# Get column names
names(river_data)
colnames(river_data) # Same as names()
# Get first and last rows
head(river_data) # First 6 rows by default
head(river_data, 3) # First 3 rows
tail(river_data) # Last 6 rows
# Summary statistics for all columns
summary(river_data)Filtering Data Frames
Filtering allows you to subset data based on conditions. Use logical operators (>, <, ==, &, |) within square brackets to select rows meeting specific criteria.
# Filter rivers with discharge > 200 m³/s
large_rivers <- river_data[river_data$Discharge_m3s > 200, ]
print(large_rivers)
# Filter by multiple conditions
# Rivers with catchment larger than 10000 km² AND discharge > 200 m³/s
large_big_rivers <- river_data[river_data$Area_km2 > 10000 &
river_data$Discharge_m3s > 200, ]
print(large_big_rivers)Adding Columns
New columns can be added to data frames using the $ operator. This example calculates specific discharge and runoff height, two key hydrological metrics for comparing catchments of different sizes.
# Calculate hydrologically relevant metrics
# 1. Specific discharge (L/s/km²) - discharge per unit area
river_data$Specific_Discharge_Ls_km2 <- (river_data$Discharge_m3s * 1000) / river_data$Area_km2
# 2. Annual runoff height (mm/year)
# This converts discharge (m³/s) to runoff depth (mm/year) over the catchment
# Formula: Q (m³/s) × seconds per year × 1000 (L/m³) / (Area in m²)
# Simplified: Q × 31,536,000 / (Area_km2 × 1,000,000) × 1000
river_data$Runoff_Height_mm <- (river_data$Discharge_m3s * 31536000) / (river_data$Area_km2 * 1e6) * 1000
print(river_data) Name Area_km2 Discharge_m3s Specific_Discharge_Ls_km2 Runoff_Height_mm
1 Enns 5915.4 191 32.28860 1018.2534
2 Mur 9769.9 138 14.12502 445.4465
3 Drau 10968.0 271 24.70824 779.1991
4 Inn 25520.0 724 28.36991 894.6734
5 Salzach 6165.4 237 38.44033 1212.2542# Display results with better formatting
cat("\n=== Austrian Rivers - Hydrological Characteristics ===\n")
=== Austrian Rivers - Hydrological Characteristics ===for (i in 1:nrow(river_data)) {
cat(sprintf("\n%s River:\n", river_data$Name[i]))
cat(sprintf(" Basin area: %s km²\n", format(river_data$Area_km2[i], big.mark = ",")))
cat(sprintf(" Mean discharge: %.1f m³/s\n", river_data$Discharge_m3s[i]))
cat(sprintf(" Specific discharge: %.1f L/s/km²\n", river_data$Specific_Discharge_Ls_km2[i]))
cat(sprintf(" Annual runoff height: %.0f mm/year\n", river_data$Runoff_Height_mm[i]))
}
Enns River:
Basin area: 5,915.4 km²
Mean discharge: 191.0 m³/s
Specific discharge: 32.3 L/s/km²
Annual runoff height: 1018 mm/year
Mur River:
Basin area: 9,769.9 km²
Mean discharge: 138.0 m³/s
Specific discharge: 14.1 L/s/km²
Annual runoff height: 445 mm/year
Drau River:
Basin area: 10,968 km²
Mean discharge: 271.0 m³/s
Specific discharge: 24.7 L/s/km²
Annual runoff height: 779 mm/year
Inn River:
Basin area: 25,520 km²
Mean discharge: 724.0 m³/s
Specific discharge: 28.4 L/s/km²
Annual runoff height: 895 mm/year
Salzach River:
Basin area: 6,165.4 km²
Mean discharge: 237.0 m³/s
Specific discharge: 38.4 L/s/km²
Annual runoff height: 1212 mm/yearCommenting Code
Code comments are explanatory text that R ignores when running your code. Good comments are essential for:
- Understanding: Remember what you did and why (future you will thank you!)
- Collaboration: Help others understand your analysis
- Documentation: Explain assumptions, data sources, and methodology
- Debugging: Temporarily disable code without deleting it
- Learning: Explain concepts to yourself as you learn
The golden rule: Write comments as if you’re explaining to someone (including yourself in 6 months) who doesn’t remember the context.
How to Comment
Use the hash symbol # for comments. Anything after # on the same line is ignored by R. Comments help explain your code logic and make it easier to understand later.
# This is a single-line comment
x <- 5 + 3 # You can also add comments at the end of a line
# This is a
# multi-line
# comment
# Calculate average daily flow in m³/s
daily_flow_data <- c(5, 6, 9, 12, 15, 10, 6, 5)
avg_flow <- mean(daily_flow_data)
# Convert flow from m³/s to L/s
flow_L_per_s <- avg_flow * 1000 # 1 m³ = 1000 LWhy Comment?
- Explanation: Describe what complex operations do
- Documentation: Provide information about inputs, outputs, data sources, or function purposes
- Debugging: Temporarily disable code without deleting it
- Readability: Break up long sections with descriptive comments
- Collaboration: Help others (or future you) understand your thought process
- Assumptions: Document important assumptions or decisions in your analysis
Good vs. Bad Comments
# BAD: States the obvious
x <- 5 # Assign 5 to x
# GOOD: Explains the meaning
catchment_area <- 648.8 # km² - Enns catchment at Schladming gauge
# BAD: Redundant with code
discharge <- discharge * 1000 # Multiply discharge by 1000
# GOOD: Explains the purpose
discharge_Ls <- discharge_m3s * 1000 # Convert from m³/s to L/s
# BAD: Vague
# Fix the data
data <- data[!is.na(data$Q), ]
# GOOD: Specific and informative
# Remove rows with missing discharge values (sensor malfunction in Jan 2020)
data <- data[!is.na(data$Q), ]Comment Styles for Different Purposes
# HEADER COMMENT: File purpose and metadata ----
# Script: Calculate annual water balance
# Author: Your Name
# Date: 2025-01-15
# Purpose: Process Schladming catchment data for 2020-2023
# Input: schladming_Q.csv, schladming_MET.csv
# Output: annual_water_balance.csv
# -----------------------------------------------
# SECTION DIVIDER (easier to spot) ================
# EXPLANATORY COMMENT: Why this approach?
# We use the Penman-Monteith equation instead of Hargreaves
# because we have full meteorological data available
# WARNING COMMENT: Important caveat
# NOTE: Data before 1981 is unreliable due to gauge relocation
# TEMPORARY COMMENT: For development
# TODO: Add error handling for missing data
# FIXME: This breaks when catchment_area = 0
# DEBUG: Print intermediate values
print(paste("Intermediate result:", temp_value))
# INLINE COMMENT: Brief clarification
n <- 0.03 # Manning's roughness coefficient for natural streamBest Practices
- Keep comments concise and clear - Be brief but informative
- Update comments when you change code - Outdated comments are worse than no comments
- Use consistent formatting - Choose a style and stick to it
- Avoid obvious comments - Don’t state what the code clearly does
- Explain “why” rather than just “what” - The code shows what, comments should explain why
- Use comments to structure code logically - Break long scripts into clear sections
- Document data sources and assumptions - Critical for reproducibility
- Comment complex calculations - Especially important for hydrological formulas
- Use comments to explain units - Very important in hydrology (m³/s vs L/s, mm vs m)
- Comment temporary workarounds - Mark things that need improvement with TODO or FIXME
Working with Data Files
In real-world applications, you’ll need to import data from external files and save results.
Reading CSV Files
CSV (Comma-Separated Values) is one of the most common formats for storing tabular data. The read.csv() function loads data into R as a data frame, with parameters to handle different delimiters and decimal separators.
# Basic CSV reading
data <- read.csv("path/to/your/file.csv")
# CSV with specific parameters (common for European data)
data <- read.csv("path/to/your/file.csv",
sep = ";", # Semicolon separator
dec = ",", # Comma as decimal point
na.strings = "-999") # How missing values are coded
# Display first few rows
head(data)
# Get summary statistics
summary(data)
# Check structure
str(data)Writing CSV Files
After processing data, you can save results to CSV files for sharing or further analysis. Set row.names = FALSE to avoid adding an extra index column.
# Save data frame to CSV
write.csv(river_data, "output/river_data.csv", row.names = FALSE)
# Or use write.table for more control
write.table(river_data,
"output/river_data.txt",
sep = ";",
dec = ",",
row.names = FALSE)Other File Formats
# Excel files (requires readxl package)
library(readxl)
data_excel <- read_excel("path/to/file.xlsx", sheet = 1)
# Save to Excel (requires writexl package)
library(writexl)
write_xlsx(river_data, "output/river_data.xlsx")Using R Libraries (Packages)
R libraries (packages) extend R’s capabilities. You need to:
- Install the package (once)
- Load the package (each R session)
# Install a package (do this once)
install.packages("readxl")
# Load the package (do this each session)
library(readxl)
# Check if package is installed, if not install it
if (!require("lubridate")) {
install.packages("lubridate")
}
library(lubridate)Data Manipulation with dplyr
The dplyr package provides intuitive functions for data manipulation. The pipe operator %>% makes code more readable by chaining operations.
The Pipe Operator (%>%)
The pipe operator passes the result from one function to the next, making code more readable by avoiding nested functions. Think of it as “then do this” in a sequence of operations.
library(dplyr)
# Without pipe (nested functions)
result1 <- round(mean(c(5, 8, 12, 15)), 2)
print(result1)
# With pipe (sequential operations)
result2 <- c(5, 8, 12, 15) %>%
mean() %>%
round(2)
print(result2)Key dplyr Functions
The dplyr package provides intuitive functions for common data manipulation tasks. Each function performs a specific operation, and they can be chained together with the pipe operator for complex workflows.
library(dplyr)
# Create example data
discharge_data <- data.frame(
Date = as.Date(c("2023-01-15", "2023-01-20", "2023-02-10",
"2023-02-25", "2023-03-05", "2023-03-15")),
Station = c("A", "A", "B", "B", "A", "B"),
Discharge = c(15.2, 18.3, 22.1, 19.5, 16.8, 20.2),
Temperature = c(5.2, 6.1, 8.3, 7.9, 9.1, 8.8)
)
# select(): Choose specific columns
discharge_data %>%
select(Date, Discharge) %>%
head()
# filter(): Keep rows that meet conditions
high_flow <- discharge_data %>%
filter(Discharge > 18)
print(high_flow)
# mutate(): Create new columns or modify existing ones
discharge_data_new <- discharge_data %>%
mutate(Discharge_Ls = Discharge * 1000,
Month = format(Date, "%B"))
print(discharge_data_new)
# arrange(): Sort rows
discharge_data %>%
arrange(desc(Discharge)) %>%
head()
# group_by() and summarise(): Calculate statistics by groups
discharge_data %>%
group_by(Station) %>%
summarise(
Mean_Discharge = mean(Discharge),
Max_Discharge = max(Discharge),
N_observations = n()
)Combining Operations
Multiple dplyr operations can be chained together to create powerful data processing workflows. This example filters, transforms, aggregates, and sorts data in a single readable sequence.
# Complex workflow with multiple operations
result <- discharge_data %>%
filter(Discharge > 16) %>% # Keep high flows
mutate(Month = format(Date, "%m")) %>% # Add month column
group_by(Station) %>% # Group by station
summarise(
Mean_Q = mean(Discharge),
Count = n()
) %>%
arrange(desc(Mean_Q)) # Sort by mean discharge
print(result)# A tibble: 2 × 3
Station Mean_Q Count
<chr> <dbl> <int>
1 B 20.6 3
2 A 17.6 2Data Visualization
Basic Plots with Base R
R has powerful built-in plotting capabilities. Base R graphics are quick to create and highly customizable, making them ideal for exploratory data analysis and hydrograph visualization.
Line Plot
Line plots are essential for visualizing time series data like discharge or precipitation over time.
# Generate example time series
dates <- seq(as.Date("2023-01-01"), as.Date("2023-12-31"), by = "month")
discharge <- c(5, 8, 15, 25, 40, 30, 20, 12, 8, 6, 4, 3)
# Create line plot
plot(dates, discharge,
type = "l", # Line type
col = "blue", # Color
lwd = 2, # Line width
main = "Monthly Discharge", # Title
xlab = "Date", # X-axis label
ylab = "Discharge (m³/s)") # Y-axis label
# Add horizontal line for mean
abline(h = mean(discharge), col = "red", lty = 2)
# Add grid
grid()
# Add legend
legend("topright",
legend = c("Discharge", "Mean"),
col = c("blue", "red"),
lty = c(1, 2),
lwd = c(2, 1))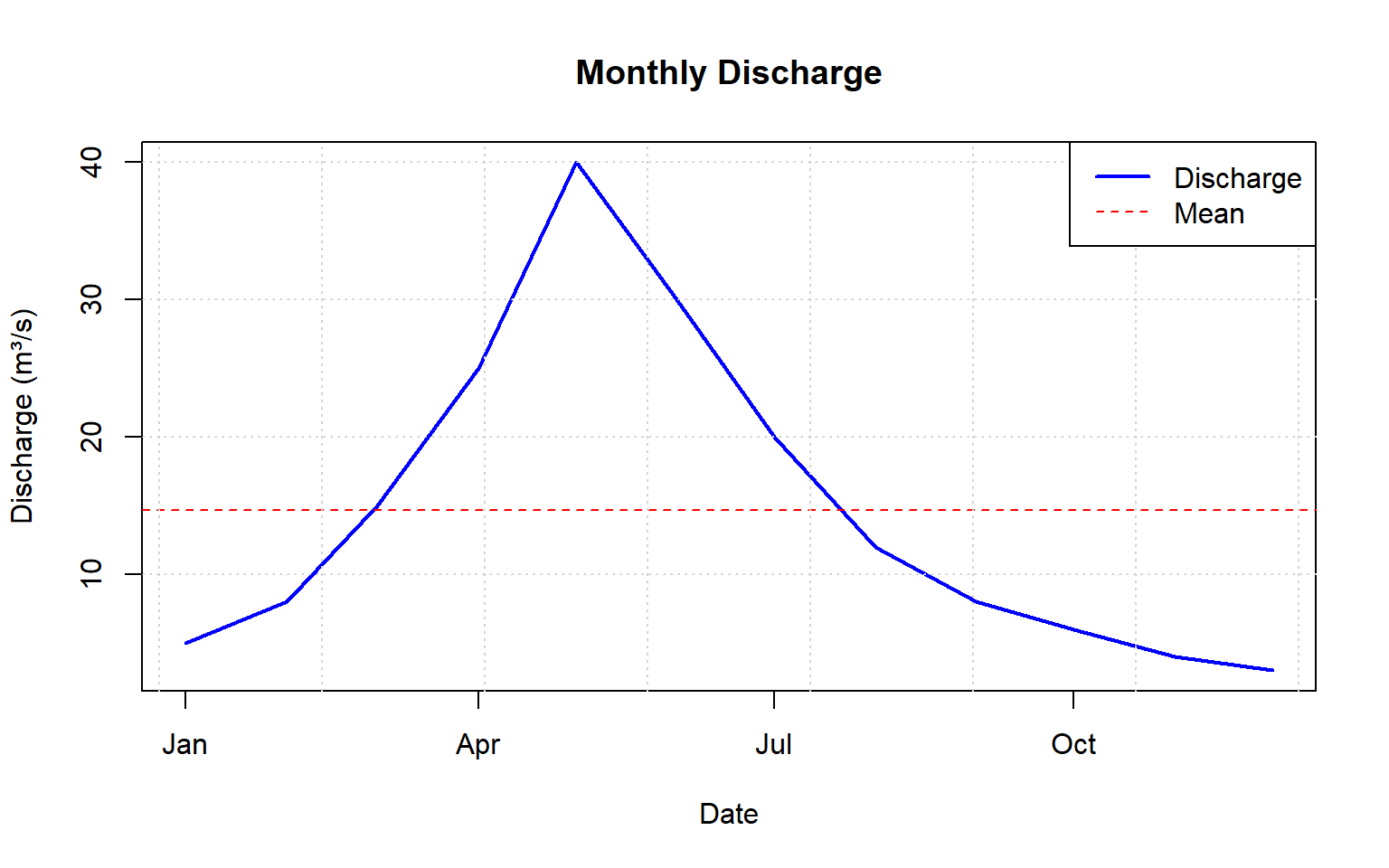
Multiple Lines
Plotting multiple data series on the same graph allows comparison between stations or variables. Add subsequent series with the lines() function after creating the initial plot.
# Create second data series
discharge2 <- discharge * 0.8 + 2
# Plot first series
plot(dates, discharge,
type = "l",
col = "blue",
lwd = 2,
ylim = range(c(discharge, discharge2)), # Set y-axis limits
main = "Discharge Comparison",
xlab = "Date",
ylab = "Discharge (m³/s)")
# Add second series
lines(dates, discharge2, col = "red", lwd = 2)
# Add legend
legend("topright",
legend = c("Station A", "Station B"),
col = c("blue", "red"),
lty = 1,
lwd = 2)
grid()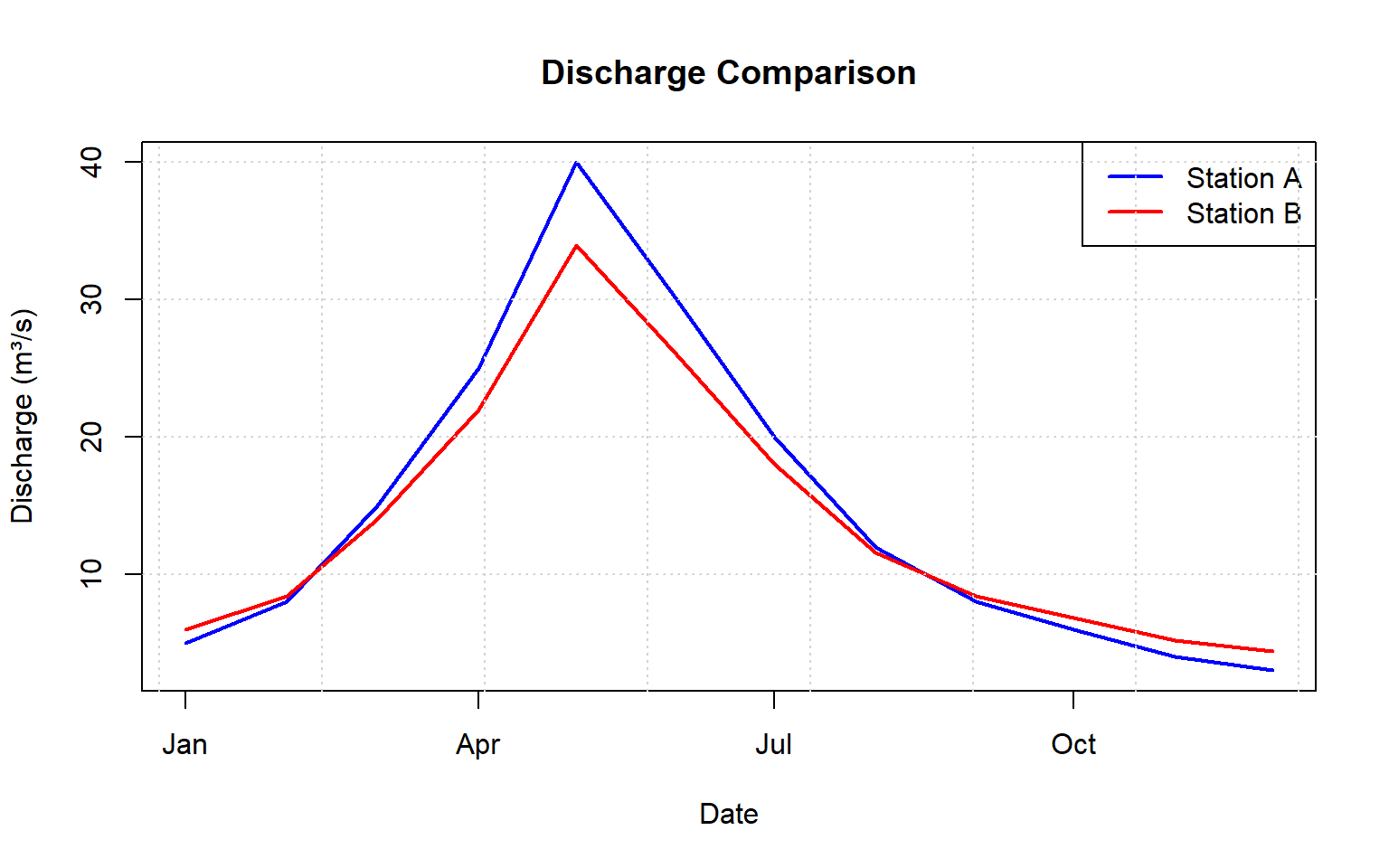
Bar Plot
Bar plots effectively display categorical data or comparisons across discrete groups. This example shows monthly rainfall totals with value labels for easy interpretation.
# Monthly rainfall data
rainfall <- c(42, 38, 52, 42, 79, 70, 78, 69, 64, 47, 46, 47)
names(rainfall) <- month.abb
# Create bar plot
barplot(rainfall,
main = "Monthly Rainfall",
xlab = "Month",
ylab = "Rainfall (mm)",
col = "skyblue",
border = "darkblue")
# Add value labels on top of bars
text(x = seq(0.7, 14.3, by = 1.2),
y = rainfall + 3,
labels = round(rainfall, 0),
cex = 0.8)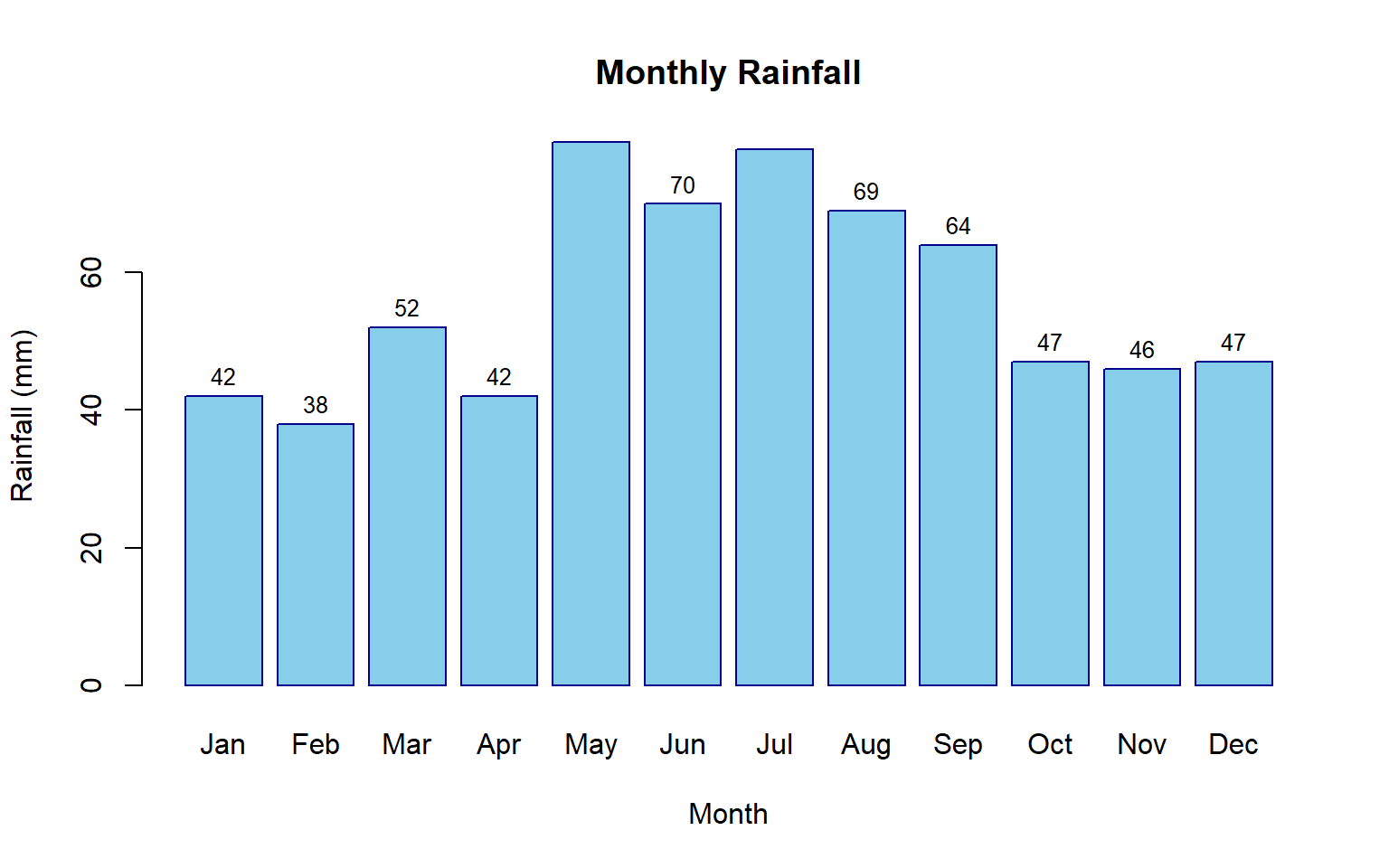
Multi-Panel Plots
The par(mfrow) function creates multiple plots in a single figure, useful for comparing different variables or showing related analyses side by side.
# Set up 2x2 panel layout
par(mfrow = c(2, 2))
# Plot 1: Line plot
plot(dates, discharge, type = "l", main = "Discharge", col = "blue")
# Plot 2: Bar plot
barplot(rainfall[1:6], main = "Rainfall (Jan-Jun)", col = "skyblue")
# Plot 3: Scatter plot
plot(discharge, rainfall, main = "Discharge vs Rainfall", pch = 16)
# Plot 4: Histogram
hist(discharge, main = "Discharge Distribution", col = "lightgreen")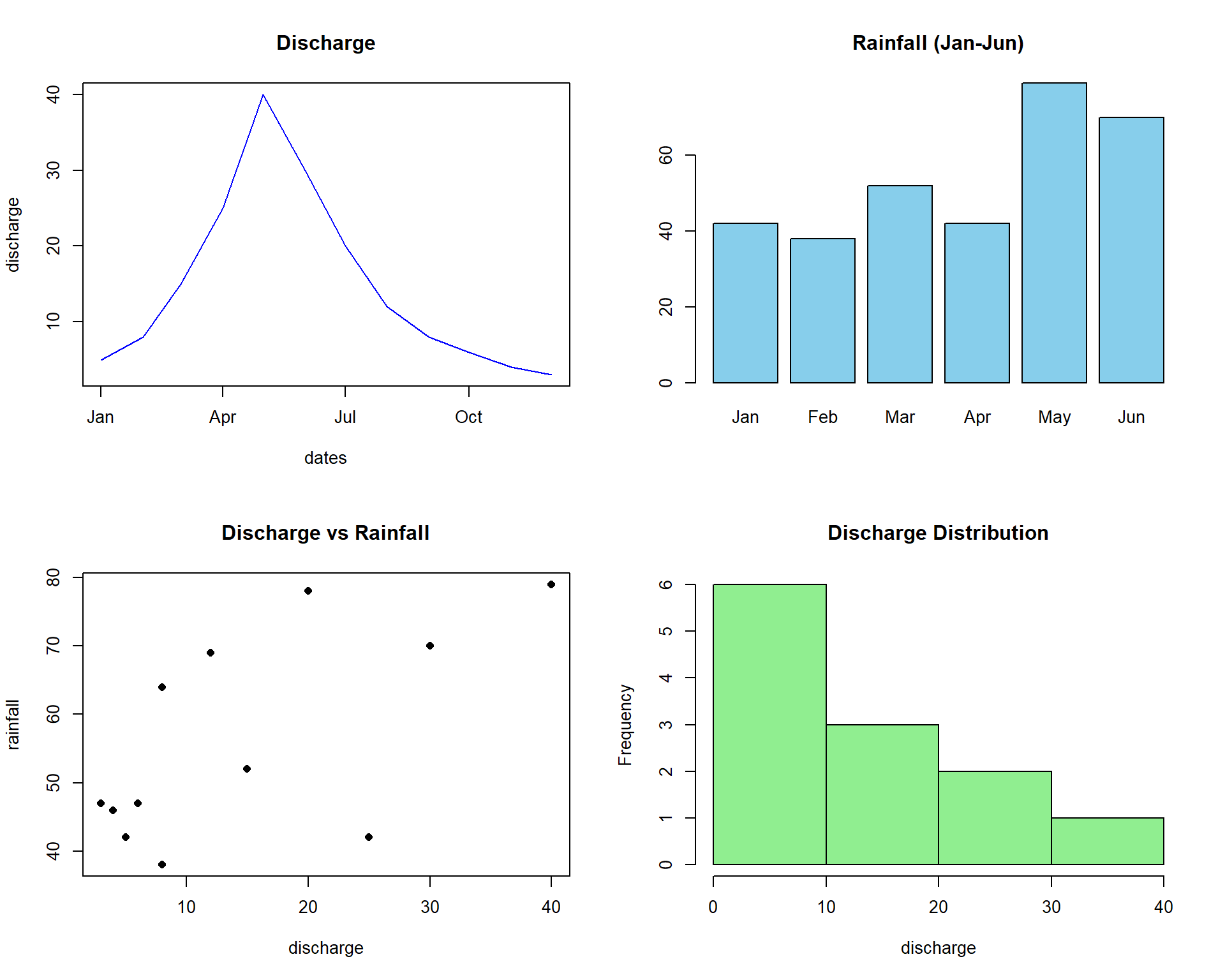
# Reset to single panel
par(mfrow = c(1, 1))Saving Plots
To save plots for reports or presentations, open a graphics device (PNG, PDF, etc.), create the plot, then close the device with dev.off(). Specify dimensions and resolution for print quality.
# Save as PNG
png("output/discharge_plot.png",
width = 15, height = 10, units = "cm", res = 300)
plot(dates, discharge, type = "l", col = "blue", lwd = 2,
main = "Monthly Discharge", xlab = "Date", ylab = "Discharge (m³/s)")
dev.off() # Close the graphics device
# Save as PDF
pdf("output/discharge_plot.pdf", width = 6, height = 4)
plot(dates, discharge, type = "l", col = "blue", lwd = 2,
main = "Monthly Discharge", xlab = "Date", ylab = "Discharge (m³/s)")
dev.off()Practical Example: Enns Catchment above Schladming
Now we’ll apply everything you’ve learned to a real-world hydrological analysis. This example uses data from the Enns catchment in the Austrian Alps.
- Station: Schladming (LamaH-CE ID: 248)
- River: Enns River
- Location: Salzburg and Styria
- Catchment area: 648.8 km²
- Elevation range: ~700-2,900 m a.s.l.
- Hydrological regime: Snowmelt-dominated alpine catchment with high flows in spring and summer and low flows in winter
What this example demonstrates:
- Reading and merging multiple data files
- Data preprocessing (dates, filtering, column renaming)
- Calculating hydrological metrics (runoff depth, water balance)
- Creating professional visualizations
- Temporal aggregation (annual and monthly)
- Saving results for further analysis
This is a somewhat complete, realistic workflow that mirrors what you’ll do in Parts II and III.
Load Required Libraries
# Uncomment the lines below to install packages (do this once)
# install.packages("dplyr")
# install.packages("lubridate")
library(dplyr)
library(lubridate)Read and Prepare Data
Proper file path configuration is essential for successful loading of data. Many errors stem from incorrect path definitions, so take time to understand and verify your paths before proceeding.
Understanding File Paths
File paths in R use forward slashes (/) or double backslashes (\\), never single backslashes (\). This differs from Windows Explorer, which displays paths with single backslashes.
Windows Explorer shows: D:\Folder\Subfolder\file.txt R requires: D:/Folder/Subfolder/file.txt or D:\\Folder\\Subfolder\\file.txt
How to Copy Paths from Windows Explorer
- Navigate to your folder in Windows Explorer
- Click in the address bar (where the path is displayed)
- Copy the path (Ctrl+C)
- In R, paste and replace all backslashes (\) with forward slashes (/)
Quick Path Fix: After pasting a Windows path in R, use Find & Replace (Ctrl+H) to replace all \ with /
# Set path to data files
# IMPORTANT: Update this path to match your local directory
path <- "D:/Lehre/HydrologieWW_II/WS25_26/example_data/"
# Read discharge data
discharge_data <- read.csv(paste0(path, "ID_248_Schladming_Q.csv"),
sep = ";", dec = ".")
# Read meteorological data
met_data <- read.csv(paste0(path, "ID_248_Schladming_MET.csv"),
sep = ";", dec = ".")
# Create date column for both datasets
discharge_data$Date <- as.Date(paste(discharge_data$YYYY,
discharge_data$MM,
discharge_data$DD, sep = "-"))
met_data$Date <- as.Date(paste(met_data$YYYY,
met_data$MM,
met_data$DD, sep = "-"))
# Merge datasets by date
hydro_data <- merge(discharge_data, met_data,
by = c("Date", "YYYY", "MM", "DD"),
all = TRUE)
# Filter to analysis period (1981-2017)
hydro_data <- hydro_data %>%
filter(YYYY >= 1981 & YYYY <= 2017)
# Display first few rows to see column names
head(hydro_data)
# Summary statistics
summary(hydro_data)Renaming Columns
When working with real-world datasets, column names are often long or contain special characters (like X2m_temp_mean from ERA5 climate data). Renaming columns to shorter, clearer names makes your code more readable.
# Method 1: Using base R names() function
# Good for renaming a single column
names(hydro_data)[names(hydro_data) == "X2m_temp_mean"] <- "temp"
# Check the result
head(hydro_data)# Method 2: Rename multiple columns at once with dplyr
hydro_data <- hydro_data %>%
rename(
temp_max = X2m_temp_max,
temp_min = X2m_temp_min,
temp_dp_max = X2m_dp_temp_max,
temp_dp_min = X2m_dp_temp_min
)
# Check all column names
names(hydro_data)Summary of methods:
- Base R
names(): Best for single column rename - dplyr
rename(): Best for multiple columns, more readable - Complete replacement:
names(data) <- c("new1", "new2", ...)replaces ALL names
Visualize Time Series
Visualizing the full time series helps identify seasonal patterns, trends, and data quality issues. This multi-panel plot shows discharge, precipitation, and temperature for the entire analysis period.
# Create multi-panel plot
par(mfrow = c(3, 1), mar = c(3, 4, 2, 2))
# Plot 1: Discharge
plot(hydro_data$Date, hydro_data$qobs,
type = "l", col = "blue",
main = "Discharge at Schladming (Enns)",
xlab = "", ylab = "Discharge (m³/s)")
abline(h = mean(hydro_data$qobs, na.rm = TRUE), col = "red", lty = 2)
# Plot 2: Precipitation
plot(hydro_data$Date, hydro_data$prec,
type = "h", col = "darkblue",
main = "Daily Precipitation",
xlab = "", ylab = "Precipitation (mm/d)")
# Plot 3: Temperature
plot(hydro_data$Date, hydro_data$temp,
type = "l", col = "red",
main = "Mean Temperature",
xlab = "Date", ylab = "Temperature (°C)")
abline(h = 0, col = "gray", lty = 2)
# Reset plot parameters
par(mfrow = c(1, 1))Calculate Annual Values
Aggregating daily data to annual values reveals long-term trends and variability. This calculation converts mean discharge to annual runoff depth for comparison with precipitation.
# Calculate catchment area
catchment_area <- 648.8 # km²
# Calculate annual runoff using dplyr
annual_runoff <- hydro_data %>%
mutate(Year = YYYY) %>%
group_by(Year) %>%
summarise(
# Mean annual discharge (m³/s)
Mean_Q = mean(qobs, na.rm = TRUE),
# Annual runoff depth (mm)
# Convert m³/s to mm/year: Q * 365.25 * 86400 * 1000 / (Area * 1e6)
Runoff_mm = Mean_Q * 365.25 * 86400 * 1000 / (catchment_area * 1e6),
# Total annual precipitation (mm)
Precip_mm = sum(prec, na.rm = TRUE),
# Mean annual temperature (°C)
Mean_Temp = mean(temp, na.rm = TRUE)
)
# Display results
print(annual_runoff)# Plot annual precipitation vs runoff
plot(annual_runoff$Precip_mm, annual_runoff$Runoff_mm,
pch = 16, col = "darkgreen",
main = "Annual Precipitation vs Runoff",
xlab = "Annual Precipitation (mm)",
ylab = "Annual Runoff (mm)")
# Add linear regression line
model <- lm(Runoff_mm ~ Precip_mm, data = annual_runoff)
abline(model, col = "red", lwd = 2)
# Add R² value
r_squared <- summary(model)$r.squared
text(min(annual_runoff$Precip_mm), max(annual_runoff$Runoff_mm),
paste("R² =", round(r_squared, 3)),
pos = 4, col = "red")
grid()Calculate Monthly Climatology
Monthly climatology shows the average seasonal cycle by aggregating all years of data into typical monthly values. This reveals the hydrological regime of the catchment.
# Calculate monthly averages across all years
monthly_clim <- hydro_data %>%
group_by(MM) %>%
summarise(
Month = first(MM),
Mean_Q = mean(qobs, na.rm = TRUE),
Mean_Precip = mean(prec, na.rm = TRUE),
Mean_Temp = mean(temp, na.rm = TRUE)
)# Create monthly climatology plot
par(mfrow = c(3, 1), mar = c(3, 4, 2, 2))
# Discharge
plot(monthly_clim$Month, monthly_clim$Mean_Q,
type = "b", col = "blue", pch = 16,
main = "Mean Monthly Discharge",
xlab = "", ylab = "Discharge (m³/s)",
xaxt = "n")
axis(1, at = 1:12, labels = month.abb)
# Precipitation
plot(monthly_clim$Month, monthly_clim$Mean_Precip,
type = "h", col = "darkblue", lwd = 3,
main = "Mean Monthly Precipitation",
xlab = "", ylab = "Precipitation (mm/d)",
xaxt = "n")
axis(1, at = 1:12, labels = month.abb)
# Temperature
plot(monthly_clim$Month, monthly_clim$Mean_Temp,
type = "b", col = "red", pch = 16,
main = "Mean Monthly Temperature",
xlab = "Month", ylab = "Temperature (°C)",
xaxt = "n")
axis(1, at = 1:12, labels = month.abb)
abline(h = 0, col = "gray", lty = 2)
par(mfrow = c(1, 1))Save Results
After analysis, save processed data and figures for documentation, reports, or further analysis. Organize outputs in a dedicated folder to maintain a clean workflow.
# Save annual data
write.csv(annual_runoff,
"D:/Lehre/HydrologieWW_II/WS25_26/example_data/outputs/schladming_annual_data.csv",
row.names = FALSE)
# Save monthly climatology
write.csv(monthly_clim,
"D:/Lehre/HydrologieWW_II/WS25_26/example_data/outputs/schladming_monthly_climatology.csv",
row.names = FALSE)
# Save a plot
png("D:/Lehre/HydrologieWW_II/WS25_26/example_data/outputs/schladming_annual_precip_runoff.png",
width = 20, height = 15, units = "cm", res = 300)
plot(annual_runoff$Precip_mm, annual_runoff$Runoff_mm,
pch = 16, col = "darkgreen", cex = 1.2,
main = "Annual Precipitation vs Runoff\nSchladming Catchment (1981-2017)",
xlab = "Annual Precipitation (mm)",
ylab = "Annual Runoff (mm)")
abline(model, col = "red", lwd = 2)
text(min(annual_runoff$Precip_mm), max(annual_runoff$Runoff_mm),
paste("R² =", round(r_squared, 3)), pos = 4, col = "red")
grid()
dev.off()Programming Essentials
This section introduces core programming concepts that will help you work more efficiently with hydrological data. While packages like dplyr handle many common tasks, understanding these fundamentals opens up new possibilities:
- Lists: Organizing complex data structures (multiple datasets, model outputs)
- For-loops: Processing multiple stations, years, or files systematically
- If-else statements: Making decisions based on data conditions
- Functions: Creating reusable tools for repeated analyses
Learning approach: These concepts can feel abstract initially. We’ve designed simple hydrological examples to illustrate each one. Focus on understanding the logic rather than memorizing every detail - these skills develop naturally with practice. Many researchers learn them gradually as specific needs arise in their work.
Lists
Lists are flexible containers that can hold different types of data - numbers, text, data frames, even other lists. Unlike vectors (which must contain the same type), lists can mix different data types and structures.
Why lists matter in hydrology:
- Store results from multiple stations or model runs
- Organize related datasets (discharge, meteorology, catchment properties)
- Return multiple values from functions
- Work with complex model outputs
# Create a list with station information
station_info <- list(
name = "Schladming",
id = 248,
area_km2 = 648.8,
elevation_m = c(700, 2900), # min and max elevation
data_years = 1981:2017
)
# Access elements by name
print(station_info$name)[1] "Schladming"print(station_info$area_km2)[1] 648.8# Access elements by position
print(station_info[[1]]) # First element[1] "Schladming"# Access nested elements
print(station_info$elevation_m[2]) # Maximum elevation[1] 2900# Check structure
str(station_info)List of 5
$ name : chr "Schladming"
$ id : num 248
$ area_km2 : num 649
$ elevation_m: num [1:2] 700 2900
$ data_years : int [1:37] 1981 1982 1983 1984 1985 1986 1987 1988 1989 1990 ...Lists for Multiple Stations
Lists excel at organizing data from multiple locations:
# Create data for multiple stations
stations <- list(
schladming = list(
id = 248,
area_km2 = 648.8,
mean_q_m3s = 18.5
),
gstatterboden = list(
id = 249,
area_km2 = 1,
mean_q_m3s = 86.2
)
)
# Access specific station data
print(stations$schladming$area_km2)[1] 648.8print(stations$gstatterboden$mean_q_m3s)[1] 86.2# Get names of all stations
names(stations)[1] "schladming" "gstatterboden"Practical tip: Lists become very useful when you start working with model outputs or processing multiple files. You’ll see them extensively in Parts II and III.
For-Loops
For-loops repeat operations for each item in a sequence. Think of them as automating repetitive tasks - instead of copying and pasting code for each station or month, write the code once and loop through all items.
When for-loops are useful in hydrology:
- Processing multiple gauging stations
- Analyzing data year-by-year or month-by-month
- Reading multiple data files
- Generating multiple plots
Basic For-Loop
# Simple loop through numbers
for (i in 1:5) {
print(paste("Iteration", i))
}[1] "Iteration 1"
[1] "Iteration 2"
[1] "Iteration 3"
[1] "Iteration 4"
[1] "Iteration 5"# Loop through a vector of names
rivers <- c("Enns", "Mur", "Drau")
for (river in rivers) {
print(paste("Processing", river, "river"))
}[1] "Processing Enns river"
[1] "Processing Mur river"
[1] "Processing Drau river"Practical Example: Monthly Aggregation
A common task in hydrology is aggregating daily data to monthly values. Here’s how a loop can help:
# Example daily discharge data for one year (365 values)
set.seed(123) # For reproducible random numbers
daily_Q <- abs(rnorm(365, mean = 15, sd = 8)) # Simulated daily discharge
days_in_year <- 1:365
# Create vector to store monthly means
monthly_Q <- numeric(12) # Pre-allocate: 12 months
# Define days per month
days_per_month <- c(31, 28, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31)
# Loop through each month
day_counter <- 1 # Track position in daily data
for (month in 1:12) {
# Define start and end day for this month
start_day <- day_counter
end_day <- day_counter + days_per_month[month] - 1
# Calculate mean discharge for this month
monthly_Q[month] <- mean(daily_Q[start_day:end_day])
# Update counter for next month
day_counter <- end_day + 1
# Print progress
cat(paste("Month", month, ": Mean Q =", round(monthly_Q[month], 2), "m³/s\n"))
}Month 1 : Mean Q = 14.79 m³/s
Month 2 : Mean Q = 16.35 m³/s
Month 3 : Mean Q = 15.47 m³/s
Month 4 : Mean Q = 14.25 m³/s
Month 5 : Mean Q = 13.87 m³/s
Month 6 : Mean Q = 15.74 m³/s
Month 7 : Mean Q = 15.48 m³/s
Month 8 : Mean Q = 14.27 m³/s
Month 9 : Mean Q = 16.02 m³/s
Month 10 : Mean Q = 16.07 m³/s
Month 11 : Mean Q = 16.72 m³/s
Month 12 : Mean Q = 15.1 m³/sProcessing Multiple Stations
Loops are invaluable when working with data from multiple locations:
# Station information
station_ids <- c(248, 249, 250)
station_names <- c("Schladming", "Gstatterboden", "Steyr")
# Create empty list to store results
results <- list()
for (i in 1:length(station_ids)) {
cat(paste("\nProcessing Station", station_ids[i], "-", station_names[i], "\n"))
# In real analysis, you would read actual data:
# data <- read.csv(paste0("station_", station_ids[i], ".csv"))
# Simulate calculating statistics
mean_Q <- runif(1, 10, 30)
max_Q <- mean_Q * runif(1, 2, 4)
# Store results in list
results[[i]] <- data.frame(
ID = station_ids[i],
Name = station_names[i],
Mean_Q = round(mean_Q, 1),
Max_Q = round(max_Q, 1)
)
}
Processing Station 248 - Schladming
Processing Station 249 - Gstatterboden
Processing Station 250 - Steyr Practical tips:
- Pre-allocate vectors: Create empty vectors before the loop for better performance
- Use meaningful counter names:
monthis clearer thaniwhen looping through months - Print progress:
cat()statements help track what’s happening in long loops - Consider alternatives: For simple operations, dplyr is often clearer than loops
If-Else Statements
If-else statements make decisions in your code based on conditions. They’re like asking questions: “If this is true, do this; otherwise, do that.”
When if-else is useful in hydrology:
- Classifying flow conditions (low/normal/high)
- Data quality checks
- Handling different seasons
- Setting warnings or alerts
Basic If-Else
# Simple if-else with river level
river_level <- 4.5 # meters
if (river_level > 5) {
print("WARNING: Severe flood risk!")
} else if (river_level > 3.5) {
print("ALERT: Moderate flood risk")
} else {
print("Normal conditions")
}[1] "ALERT: Moderate flood risk"Classifying Discharge
A common application is classifying flow conditions (or some other variable):
# Function to classify single discharge value
classify_flow <- function(discharge) {
if (discharge < 5) {
return("Low flow")
} else if (discharge < 15) {
return("Normal flow")
} else {
return("High flow")
}
}
# Test with different values
cat("Q = 3 m³/s:", classify_flow(3), "\n")Q = 3 m³/s: Low flow cat("Q = 10 m³/s:", classify_flow(10), "\n")Q = 10 m³/s: Normal flow cat("Q = 20 m³/s:", classify_flow(20), "\n")Q = 20 m³/s: High flow Vectorized If-Else with ifelse()
When working with multiple values, use ifelse() which is faster:
# Classify multiple discharge values at once
discharge_values <- c(3, 10, 20, 8, 25, 2)
# ifelse() works on entire vectors
flow_class <- ifelse(discharge_values < 5, "Low",
ifelse(discharge_values < 15, "Normal", "High"))
# View results
result <- data.frame(
Discharge_m3s = discharge_values,
Classification = flow_class
)
print(result) Discharge_m3s Classification
1 3 Low
2 10 Normal
3 20 High
4 8 Normal
5 25 High
6 2 LowKey difference: Use if for single values, use ifelse() for vectors (multiple values at once).
Creating Functions
Functions are reusable pieces of code that take inputs, perform calculations, and return results. Think of them as creating your own custom tools.
When functions are useful in hydrology:
- Repeating the same calculation for multiple stations
- Implementing hydrological equations (Manning’s equation, rating curves, etc.)
- Ensuring consistency in calculations
- Making code more readable and organized
Basic Function Structure
# Simple function to convert discharge units
convert_Q_to_Ls <- function(Q_m3s) {
# Convert m³/s to L/s
Q_Ls <- Q_m3s * 1000
return(Q_Ls)
}
# Use the function
discharge <- 5.3 # m³/s
discharge_Ls <- convert_Q_to_Ls(discharge)
cat("Discharge:", discharge, "m³/s =", discharge_Ls, "L/s\n")Discharge: 5.3 m³/s = 5300 L/sFunction with Multiple Inputs
# Calculate runoff coefficient
# (ratio of runoff to precipitation)
calculate_runoff_coeff <- function(precip_mm, runoff_mm) {
# Input: precipitation and runoff in mm
# Output: runoff coefficient (0-1)
coeff <- runoff_mm / precip_mm
return(coeff)
}
# Test the function
annual_precip <- 1200 # mm
annual_runoff <- 400 # mm
rc <- calculate_runoff_coeff(annual_precip, annual_runoff)
cat("Runoff coefficient:", round(rc, 3), "\n")Runoff coefficient: 0.333 Function Returning Multiple Values
Functions can return multiple results using a list:
# Calculate water balance components
water_balance <- function(precip_mm, runoff_mm) {
# Calculate water balance components
# Assuming evapotranspiration = precipitation - runoff
et_mm <- precip_mm - runoff_mm
runoff_coeff <- runoff_mm / precip_mm
# Return multiple values as a list
results <- list(
evapotranspiration_mm = et_mm,
runoff_coefficient = runoff_coeff,
water_balance_closed = (precip_mm == runoff_mm + et_mm)
)
return(results)
}
# Use the function
wb <- water_balance(1200, 400)
cat("Evapotranspiration:", wb$evapotranspiration_mm, "mm\n")Evapotranspiration: 800 mmcat("Runoff coefficient:", round(wb$runoff_coefficient, 3), "\n")Runoff coefficient: 0.333 cat("Balance closed:", wb$water_balance_closed, "\n")Balance closed: TRUE Practical Example: Rating Curve
Rating curves convert water stage (height) to discharge - a fundamental tool in hydrology:
# Rating curve: Q = a * h^b
rating_curve <- function(stage_m, a = 10, b = 2.5) {
# Calculate discharge from water stage
# Using power law: Q = a * h^b
#
# Input:
# stage_m: water stage in meters
# a, b: rating curve parameters (with default values)
#
# Output: discharge in m³/s
discharge <- a * stage_m^b
return(discharge)
}
# Test with different stage values
stages <- c(0.5, 1.0, 1.5, 2.0, 2.5)
discharges <- rating_curve(stages)
# Display results
rating_table <- data.frame(
Stage_m = stages,
Discharge_m3s = round(discharges, 2)
)
print(rating_table) Stage_m Discharge_m3s
1 0.5 1.77
2 1.0 10.00
3 1.5 27.56
4 2.0 56.57
5 2.5 98.82# Plot the rating curve
plot(stages, discharges,
type = "b", pch = 16, col = "blue",
main = "Rating Curve",
xlab = "Stage (m)",
ylab = "Discharge (m³/s)")
grid()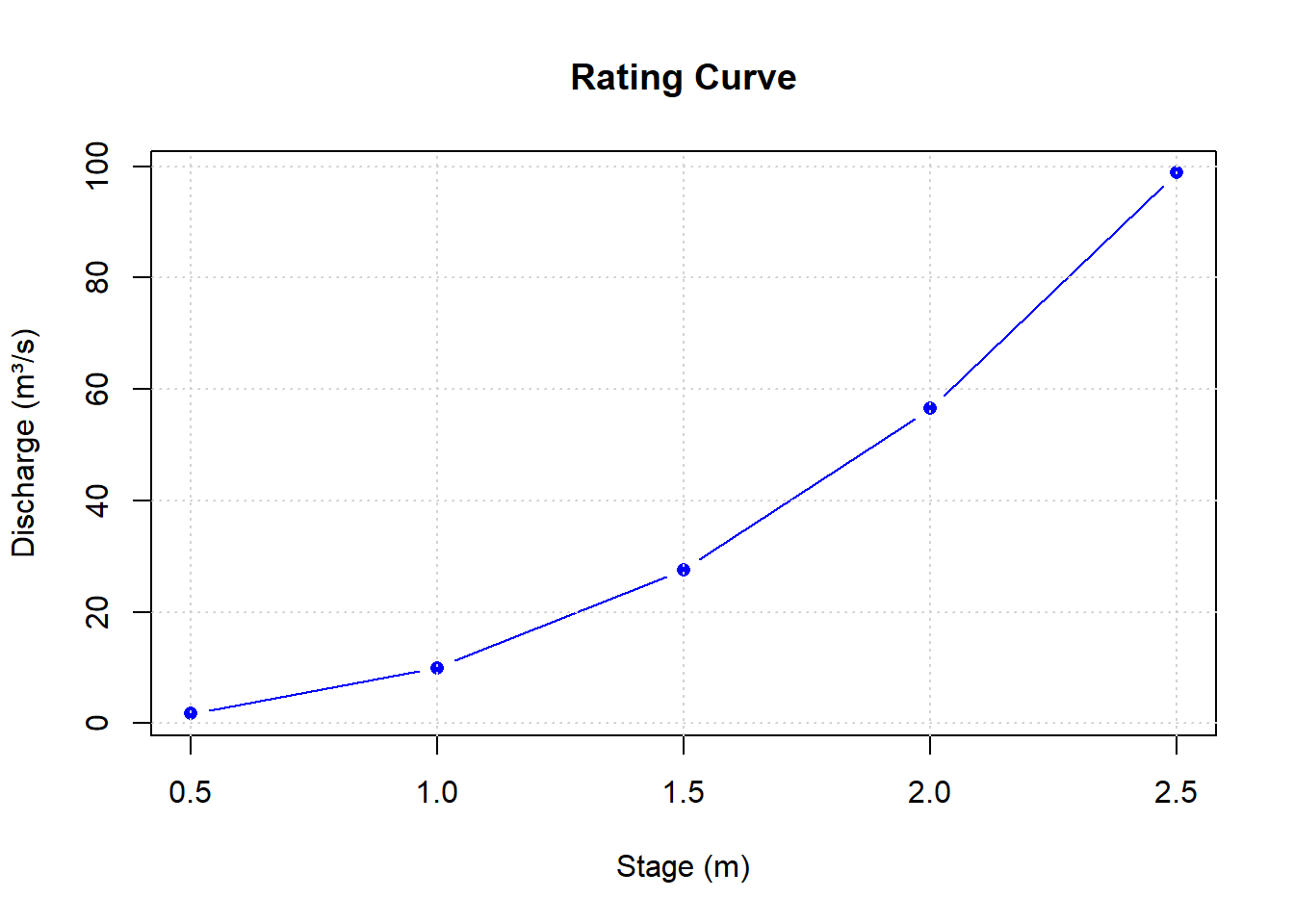
Function tips:
- Choose clear names:
calculate_runoff_coeff()is better thancalc_rc() - Add comments: Explain what the function does, especially the inputs and outputs
- Test your functions: Try different inputs to make sure they work as expected
- Use default values: For parameters that usually stay the same (like
aandbin rating curves)
Summary and Next Steps
What You Learned
In this introduction to R and RStudio, you learned:
- R Basics: Variables, data types, basic operations
- Data Structures: Vectors and data frames
- Data Management: Reading, writing, and manipulating data
- Data Visualization: Creating plots with base R
- Data Manipulation: Using dplyr and the pipe operator
- Practical Application: Analyzing Schladming catchment data
- Getting Help: Using built-in help, online resources, and LLMs effectively
Skills for Parts II and III
You now have the foundation needed for:
- Part II: GR4J Modeling
- Reading meteorological and discharge data
- Data preprocessing and merging
- Running hydrological models
- Analyzing model outputs
- Part III: Hydrological Analysis
- Time series analysis
- Statistical calculations
- Advanced visualizations
- Trend analysis
Additional Resources
General R Resources
- R for Data Science: https://r4ds.had.co.nz/
- RStudio Cheatsheets: https://www.rstudio.com/resources/cheatsheets/
Hydrology-Specific Resources
- CRAN Task View: Hydrology: https://cran.r-project.org/web/views/Hydrology.html
- Slater, L. J., et al. (2019): Using R in hydrology: a review of recent developments and future directions. Hydrology and Earth System Sciences, 23, 2939–2963. https://doi.org/10.5194/hess-23-2939-2019
- Delaigue, O., et al. (2023): airGRteaching: an open-source tool for teaching hydrological modeling with R. Hydrology and Earth System Sciences, 27, 3293–3327. https://doi.org/10.5194/hess-27-3293-2023
Tips for Success
- Practice regularly: The best way to learn programming is by doing
- Start small: Begin with simple tasks and gradually increase complexity
- Use comments: Document your code for future reference
- Ask for help: Use built-in help, online resources, and LLMs
- Learn from errors: Debugging is a normal part of programming
- Stay organized: Use meaningful variable names and file structures
Good luck with your hydrological modeling journey!
Document created: 2026-03-16
Comment Hierarchies
Using hierarchical comments helps organize code into logical sections:
In RStudio, press
Ctrl+Shift+O(Windows/Linux) orCmd+Shift+O(Mac) to display the document outline based on your comment hierarchy. Adding####after section headings makes them appear in the outline.